Blogs
Strand’s Approach to Delivering Scalable, Clinical-Grade AI Infrastructure for Digital Histopathology
In many diseases, histopathology, which involves the microscopic examination of tissues and cells, is widely used for diagnosis, particularly in cancer. It plays a critical role in identifying, classifying, and characterizing different tissue types, cell populations, and disease states. Traditionally, this assessment relies on manual interpretation of H&E-stained slides, where pathologists examine tissue morphology based on their expertise and experience. While essential, manual analysis can be time-consuming and may show variability due to tissue complexity and differences in observer interpretation.
With the increasing digitization of histopathology images, computational methods are being explored to support analysis. Deep learning models can assist by automating image classification, helping improve consistency and enabling more scalable workflows. AI is increasingly being explored as a supportive tool in digital histopathology to augment diagnostic workflows.
However, applying AI in digital histopathology within real-world research and clinical environments depends on structured data pipelines, validation discipline, reproducibility, and governance. At Strand, we design and implement scalable, end-to-end AI workflows that transform fragmented imaging data into validated, interpretable, and deployable systems.
Infrastructure Supporting AI in Digital Histopathology
Our cloud-optimized infrastructure supports clinical-grade workflows while enabling interactive visualization, downloadable results, and real-time quality control dashboards.
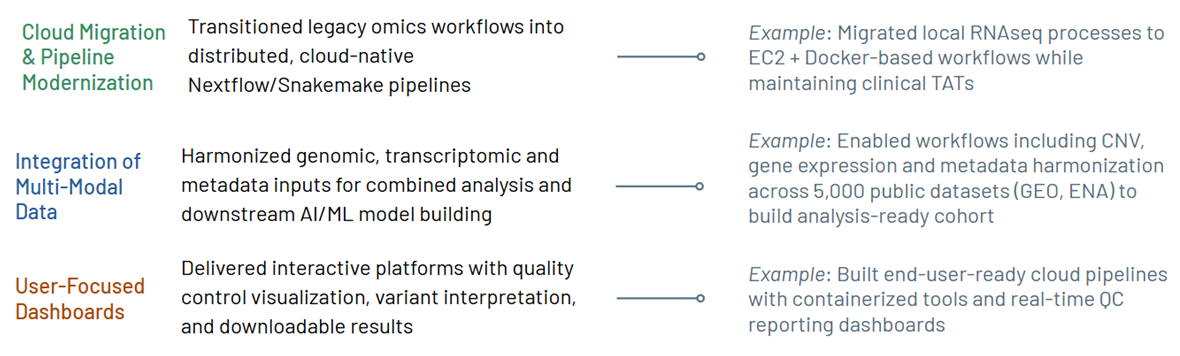
What we offer:
- Cloud-native pipelines
- FAIR data structuring
- Multi-modal integration
- Experiment tracking
- Reproducibility
- Interpretability
- Clinical-grade validation
Case Study: Breast Cancer Histopathology Classification
To demonstrate our approach in practice, we applied it to breast cancer histopathology classification. We used two widely recognized benchmark datasets:
- BACH: dataset of 400 images across four classes at a fixed magnification
- Image size 2048 x 1536 pixels
- Magnification: 200x
- BreakHis: dataset of 7,909 images from 82 patients, spanning multiple magnifications
- Benign: 4 subclasses
- Malignant: 4 subclasses
- Image size: 700 X 460 pixels
- Magnifications: 40x, 100x, 200x, 400x
Using both datasets enabled us to evaluate performance across structured and heterogeneous imaging environments.
In both datasets, combining two deep learning models improved performance compared to using a single model.
- On BACH, Ensemble (ResNet50, VGG16) accuracy reached 80.67%.
- On BreakHis, accuracy reached 86.18%, with improved sensitivity across several subclasses.
All results were validated using cross-validation and reported with 95% confidence intervals to ensure reliability. This demonstrates not only model performance, but a reproducible and validated AI workflow capable of operating across datasets of varying complexity.
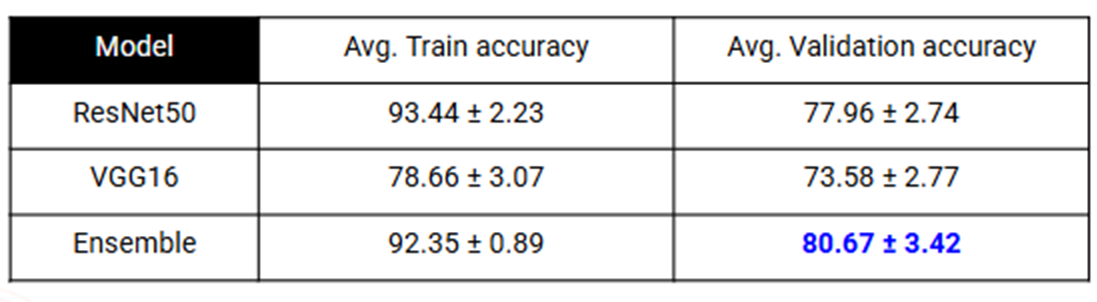
Results: BACH
Patch-wise Accuracy: We report the 95% confidence interval for the average accuracy across all 5 folds
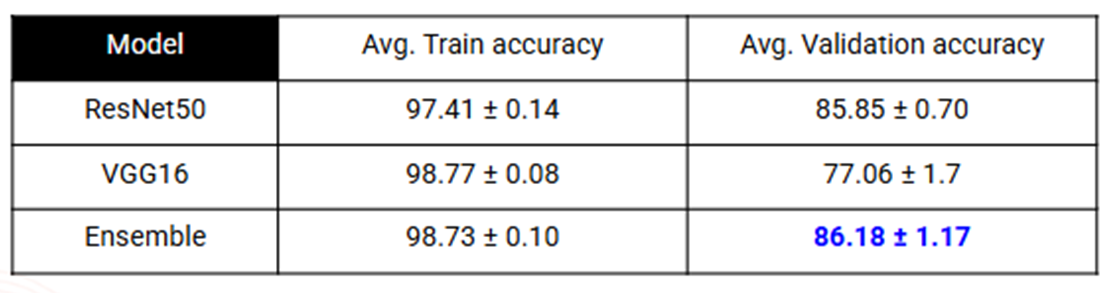
Results: BreakHis
Image-wise Accuracy: We report the 95% confidence interval for the average accuracy across all 5 folds
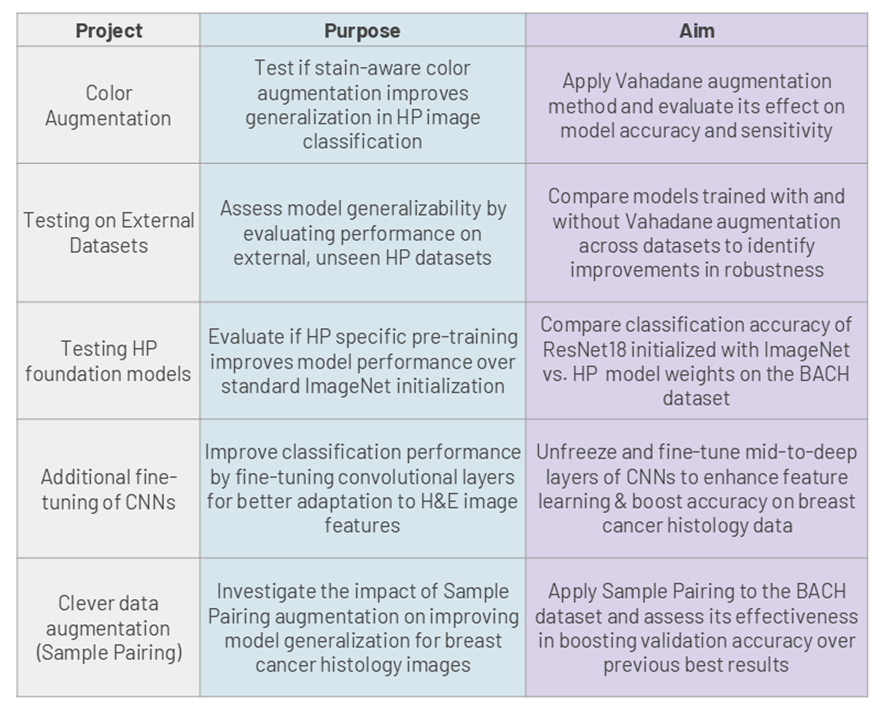
The study is being extended to additional applications as ongoing projects, as shown in the table below:
If you are looking to implement scalable, governed AI workflows in digital histopathology or integrate imaging data into a unified architecture, you can reach out to us at [email protected].
References:
- Greeley C, Holder L, Nilsson EE, Skinner MK. Scalable deep learning artificial intelligence histopathology slide analysis and validation. Sci Rep. 2024.
- McCaffrey C, Jahangir C, Murphy C, Burke C, Gallagher WM, Rahman A. Artificial intelligence in digital histopathology for predicting patient prognosis and treatment efficacy in breast cancer. Expert Rev Mol Diagn. 2024.
- Mezei T, Kolcsár M, Joó A, Gurzu S. Image Analysis in Histopathology and Cytopathology: From Early Days to Current Perspectives. J Imaging. 2024.
- Tiwari A, Ghose A, Hasanova M, et al. The current landscape of artificial intelligence in computational histopathology for cancer diagnosis. Discov Oncol. 2025.

Data Harmonization Series
22 Mar 2024
Resolving Ontology Inconsistencies: Insights from Strand's Approach

Precision Medicine
30 Jun 2025
Strand’s Bioinformatics Expertise Enables Liquid Biopsy Assay Development for NSCLC with 70% Cost Reduction and 2–5% VAF Sensitivity

Precision Medicine
26 Jun 2025
Strand’s qPCR Reporting Platform Accelerates Infectious Disease Diagnosis With ~2 min Turnaround Times
Let's Connect
Let's Connect

download the case study.




